« Previous « Start
7 GENO Algorithmic Details
7.1 Introduction
The Genetic Algorithm (or GA for short) is a recent development in
the arena of numerical search methods. GAs belong to a class of
techniques called
Evolutionary Algorithms, including
Evolutionary Strategies, Evolutionary Programming and Genetic
Programming. One description of GAs is that they are stochastic
search procedures that operate a population of entities; these
entities are suitably coded candidate solutions to a specific
problem; and the GA solves the problem by artificially mimicking
evolutionary processes observed in natural genetics.
Naturally, terminology from the field of natural genetics is used to
describe genetic algorithms. In a biological organism, the structure
that encodes how the organism is to be constructed is called the
chromosome. One or more chromosomes may be required to
completely specify the organism; the complete set of chromosomes is
called a
genotype, and the resulting organism is called a
phenotype. Each chromosome comprises a set of genes, each
with a specific position or locus on the chromosome. The loci in
conjunction with the values of the genes (which are called the
alleles) determine what characteristics are expressed by the
organism.
In the GA analogy, a problem would first be coded and in this
regard, genetic algorithms have traditionally used a binary
representation in which each candidate solution is coded as a string
of 0's and 1's. The GA's "chromosomes" are therefore the strings of
0's and 1's, each representing a different point in the space of
solutions. Normally each candidate solution (or "organism") would
have only one chromosome, and so the terms
organism,
chromosome and
genotype are often used synonymously in
GA literature. At each position on a chromosome is a
gene
that can take on the
alleles 0 or 1; the phenotype is the
decoded value of the chromosome. The population of candidate
solutions represent a sample of different points of the search
space, and the algorithm's genetic processes (see below) are such
that the chromosomes evolve and become better and better
approximations of the problem's solution over time.
Before a GA can be run, a
fitness function is required: this
assigns a figure of relative merit to each potential solution.
Before fitness values can be assigned, each coded solution has to be
decoded and evaluated, and the module designed to do this is
generally called the
evaluation function. But whereas the
evaluation function is a problem-specific mapping used to provide a
measure of how individuals have performed in the problem domain, the
fitness function, on the other hand, is a problem-independent
mapping that transforms evaluation values into an allocation of
reproductive opportunities. For any given problem therefore,
different fitness functions may be defined. At each generation, the
chromosomes in the current population are rated for their
effectiveness as candidate solutions, and a process that emulates
nature's survival-of-fittest principle is applied to generate a new
population which is then "evolved" using some genetic operators
defined on the population. This process is repeated a sufficient
number of times until a good-enough solution emerges. The three
primary genetic operators focused on in practice are
selection,
crossover and
mutation.
7.2 Selection
This operator is sometimes called
Reproduction. The
reproduction operation is in fact comprised of two phases: the
selection mechanism and the sampling algorithm. The selection
mechanism assigns to each individual x, a positive real number,
called the
target sampling rate (or simply: fitness), which
indicates the
expected number of offspring reproduced by x at
generation t. In the commonly used
fitness proportionate
selection method, an individual is assigned a target sampling rate
equal to the ratio of the individual's evaluation to the average
evaluation of all chromosomes in the population. This simple scheme
however suffers from the so-called scaling problem where a mere
shift of the underlying function can result in significantly
different fitness values. A technique that has been suggested to
overcome this is to assign target sampling rates according to some
form of population ranking scheme. Here, individuals are first
assigned a rank based on their performance as determined by the
evaluation function; thereafter, the sampling rates are computed as
some linear or non-linear function of the ranks.
After fitness values have been assigned, the sampling algorithm then
reproduces copies of individuals to form an intermediate mating
population. The most common method of sampling the population is by
the
roulette wheel method in which each individual is
assigned a slice of a virtual roulette wheel which is proportional
to the individual's fitness. To reproduce a population of size P,
the wheel is spun P times. On each spin, the individual under the
wheel's maker is selected to be in the mating pool of parents who
are to undergo further genetic action. An alternative approach and
one which minimizes
spread is Stochastic Universal Sampling
(SUS). Here, rather than spin the wheel P times to select P parents,
SUS spins the wheel once but with P equally spaced pointers which
are used to select the P parents simultaneously. Reproduction may
also be done by a tournament selection. A typical implementation is
as follows. Two individuals are chosen at random from the population
and a random number r is chosen between 0 and 1. If r < k (where k
is a tuning parameter, say 0.75), the fitter of the two individuals
is selected to go into the mating pool; otherwise the less fit
individual is chosen. The two are then returned to the original
population and can be selected again.
Reproductive processes may be implemented in
generational or
steady-state mode. Generational reproduction replaces the
entire population with a new population, and the GA is said to have
a generation gap of 1. Steady-state reproduction on the other hand
replaces only a few individuals at a time.
Elitism is an
addition to selection that forces the GA to retain some number of
the best individuals at each generation. Such individuals can be
lost if they are not selected to reproduce or if they are operated
on by the genetic operators.
The selection operator is the driving force in GAs, and the
selection pressure is a critical parameter. Too much
selection pressure may cause the GA to converge prematurely; too
little pressure makes the GA's progress towards the solution
unnecessarily slow.
7.3 Crossover
The crossover operation is also called recombination. It is
generally considered to be the main exploratory device of genetic
algorithms. This operator manipulates a pair of individuals (called
parents) to produce two new individuals (called offspring or
children) by exchanging corresponding segments from the parents'
coding. The simplest form of this operator is the single-point
crossover, and this is as illustrated below where the crossover
point is the position marked by the symbol, |.
Parent1: (a b c | d e) Cross over at position 3 Child1: (a b c | 4 5)
-------------------------->
Parent2: (1 2 3 | 4 5) Child2: (1 2 3 | d e)
Other binary-coded crossover operators which are variations of the
above scheme have since been defined, e.g., two-point crossover,
uniform crossover and shuffle crossover. For real-coded GAs,
recombination is usually defined in a slightly different way. We
mention three crossover operators that are employed by GENO:
-
ARITHMETIC CROSSOVER. This operator produces two offspring that are convex
combinations of the parents. If the chromosomes ckv = (xk1, xk2,...,xkN) and ckw = (xk1, xk2,...,xkN) are selected for crossover,
the offspring are defined as:
ck1 = α *ckv+(1−α)*ckw and ck2 =
α *ckw+(1−α)*ckv, where α 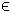 [x_L,
x_U]
[x_L,
x_U]
- HEURISTIC CROSSOVER. This operator combines two chromosomes and produces
one offspring as follows: if ckv and ckw are two parent chromosomes such that
the fitness of ckv is not worse than that of ckw, then the offspring
is:
ckx = ckv+α *(ckv−ckw), where α 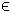 [x_L,
x_U]
[x_L,
x_U]
Here, the idea is to use the "quasi-gradient" of the evaluation function as a means of directing the search
process.
- DIFFERENTIAL CROSSOVER. This operator uses three parents: one parent is taken as
the "base", and the other two are used to generate the search direction.
Thus, if uTB, uTV and uTW, are the parent chromosomes with uTB as the "base", then the
offspring are:
uT1 = uTB+α *(uTW−uTV) and uT2 = uTB+α *(uTV−uTW),
where α 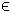 [x_L,x_U]
[x_L,x_U]
In GENO, the factor α is pre-selected from the unit interval, although random variable may also be
used.
7.4 Mutation
By modifying one or more of the gene values of an existing
individual, mutation creates new individuals and thus increases the
variability of the population. The mutation operator ensures that
the probability of reaching any point in the search space is never
zero. The mutation operator is applied to each gene of the
chromosome depending on whether a random deviate drawn from a
certain probability distribution is above a certain threshold.
Usually the uniform or normal distribution is assumed. Again,
depending on the representation adopted, variations of the basic
mutation operator may be defined.
7.5 Why Genetic Algorithms Work
Currently, there are several competing theories that attempt to
explain the macroscopic behavior of GAs. The original description of
GAs as
schema processing algorithms by John Holland (1975)
has underpinned most of the theoretical results derived to date.
However, other descriptive models based on Equivalence Relations,
Walsh functions, Markov Chains and Statistical Mechanics have since
been developed. A survey of these models is beyond the scope of this
introductory exposition. Instead, we provide a sketch of the logic
leading up to one of the main explanatory models, namely
The
Building Block Hypothesis. We begin by stating some definitions.
-
DEFINITION A1: [Schema; Schemata] A schema is a template that
defines the similarities among chromosomes which is built by
introducing the don't care symbol (*) in the alphabet of genes.
It represents all chromosomes which match it at every locus
other than the '*' positions. For example, the schema (1 0 * * 1)
represents of four chromosomes, i.e.: (1 0 1 1 1), (1 0 1 0 1),
(1 0 0 1 1) and (1 0 0 0 1). A collection of schema is a schemata.
- DEFINITION A2: [Order] The order of a schema S (denoted by o(S)) is
the number of non-don't care positions in the schema.
For example, the schemata S1 = (1 0 * * 1), S2= (* 0 * * 1), S3= (* * 1 * *)
are of orders 3, 2 and 1, respectively.
- DEFINITION A3: [Defining Length] The defining length of a schema S
(denoted by δ(S) ) is the positional distance between the first and last
fixed positions (i.e., the non-don't care sites) in the schema. It defines the
compactness of the information contained in the schema. For example,
defining lengths of the three schemata S1 = (1 0 * * 1), S2= (* 0 * * 1),
S3= (* * 1 * *) are δ(S1) = 4, δ(S2) = 3 and δ(S3) = 0, respectively.
- DEFINITION A4: [Schema Fitness] The schema fitness is the average of the fitness
of all chromosomes matched by the schema in a given population. That is, given
the evaluation function eval(.) defined on a population of chromosomes xj
of size P, the fitness of schema S at generation t is:
| |
eval(S,t) = |
|
eval(xj/P) (3) |
The evolutionary process of GAs consists of four basic steps which
are consecutively repeated, namely:
t <- t+1
select P(t) from P(t-1);
recombine and mutate P(t);
evaluate P(t)
The main evolutionary process takes place in the
select,
recombine and
mutate phases. After the selection step,
we can expect to have ξ(
S,
t+1) chromosomes matched by the schema
S in the mating pool. For an average chromosome matched by the
schema S, the probability of its selection is equal to
eval(S,t)/F(t) where F(t) is the total fitness for the whole
population. Since the number of chromosomes matched by schema S
before selection is ξ(
S,
t), and the number of single chromosome
selections is P, it follows that:
ξ(
S,
t+1)=ξ(
S,
t)*
P*
eval(
S,
t)/
F(
t) (4)
or, in terms of the average fitness of the population,
F(
t):
ξ(
S,
t+1)=ξ(
S,
t)*
eval(
S,
t)/
F(
t) (5)
In other words, the number of chromosomes grows as the ratio of the
fitness of the schema to the average fitness of the population. This
means that an above-average schema receives an increasing number of
matching chromosomes in the next generation, a below-average schema
receives a decreasing number of chromosomes, and an average schema
remains the same.
The schema growth equation
5 however has to be modified
to take into account the destructive effects of recombination and
mutation. For chromosomes of length m, the crossover site is
selected uniformly from among (m - 1) possible sites. A schema S
would be destroyed if the crossover site is located within its
defining length. The probability of this happening is
pd(
S)=δ(
S)/(
m−1) and, hence, the probability of a schema
surviving the crossover operation is,
The crossover operator is however only applied selectively according
to some probability (
ps, say). Furthermore, even when the
crossover site is within the defining length there is always a
finite chance that the schema may survive. These considerations
dictate the modification of
6 to,
ps(
S) ≥ 1−
pc*(δ(
S)/(
m−1)) (7)
Thus, the combined effects of
selection and
recombination are summarized by:
ξ(
S,
t+1) ≥ ξ(
S,
t)*(
eval(
S,
t)/
F(
t))*(1−
pc*(δ(
S)/(
m−1))) (8)
The mutation operator changes a single gene with probability
pm. It is clear that
all the fixed positions of a schema
must remain intact if the schema is to survive mutation. Since each
mutation is independent of all others, the probability of a schema S
surviving mutation is therefore:
And for
pm ≪ 1,
ps(
S) may be approximated by the first
two terms of its binomial expansion, i.e.:
Therefore, ignoring higher-order terms involving products of
pc
and
pm, the combined effects of selection, crossover and
mutation is summarized by the following reproductive schema growth
inequality:
ξ(
S,
t+1) ≥ ξ(
S,
t)*(
eval(
S,
t)/
F(
t))*(1−
pm*
o(
S)−
pc*(δ(
S)/(
m−1))) (11)
Clearly, the disruptive effects of mutation and crossover are
greater, the higher the order, and the longer the defining length of
the schema. One can therefore expect that later generations of
chromosomes would increasingly be comprised of short, low-order
schemata of above-average fitness. This observation is captured by
the
Schema Theorem which states:
- THEOREM A1 [Schema Theorem] Short, low-order, above-average schemata receive exponentially increasing trials in
subsequent generations of a genetic algorithm.
An immediate result of this theorem is the assertion that genetic
algorithms explore the search space by short, low-order schemata
which are used for information exchange during recombination. This
observation is expressed by the Building Block Hypothesis which
states:
- HYPOTHESIS A1 [Building Block Hypothesis] A genetic algorithm seeks near-optimal performance through the
juxtaposition of short, low-order, high-performance schemata called building blocks.
Over the years, many GA applications which support the building
block hypothesis have been developed in many different problem
domains. However, despite this apparent explanatory power, the
hypothesis is not universally valid. In particular, it is easily
violated in the so-called deceptive problems.
7.6 Setting GA Parameters
Before one can use a GA, one needs to specify some parameter values
namely the selection pressure, the population size, and the
crossover and mutation rates. Both theoretical and empirical studies
show that "optimal" values for these parameters depend on how
difficult the problem at hand is. And since prior determination of
the degree of difficulty a particular problem poses is hard, there
are no generally accepted recipes for choosing effective parameter
values in every case. However, many researchers have developed good
heuristics for these choices on a variety of problems, and this
section outlines some their recommendations.
7.6.1 Experimental Studies
De Jong (1975)
Kenneth A. De Jong tested various combinations of GA parameters on
five functions with various characteristics including continuous and
discontinuous, convex and non-convex, uni-modal and multi-modal,
deterministic and noisy for his PhD Thesis. His function suite has
since been adopted by many researchers as the standard test bed for
assessing GA designs.
De Jong used a simple GA with roulette wheel selection, one-point
cross-over and simple mutation to investigate the effects of four
parameters namely: population size, crossover rate, mutation rate
and the generation gap. His main conclusions were that:
- Increasing the population size resulted in better long-term
performance, but smaller population sizes responded faster and
therefore exhibited better initial performance.
- Mutation is necessary to restore lost alleles but this
should be kept low at a low rate for otherwise the GA degenerates
into a random search.
- A cross-over probability of around 0.6 worked best.
But increasing the number of cross-over points degraded performance.
- A non-overlapping population model worked better in general.
- In summary, he concluded that the following set of parameters
were efficient (at least for the functions that he studied):
population size - 50 - 100; crossover probability - 0.6; mutation
probability - 0.001; generation gap - 1.
De Jong's work was very important in that it provided practical
guidelines for subsequent applications of GA's. His recommendations
for the various parameters have been so widely adopted that they are
sometimes referred to as "the standard settings". But subsequence
research revealed that applying De Jong's parameter values cases can
be a serious mistake in some cases.
Schaffer, Caruana, Eshelman and Das (1989)
Recognizing that parameter values can have a significant impact on
the performance of a GA and that a more thorough investigation was
needed, Schaffer et al. (1989) expanded De Jong's experimental
setup. In addition to the five functions that he had studied, they
introduced five more and performed a more exhaustive assessment of
the direct and cross effects of the various parameters on a GA's
performance. A notable observation they made was that good GA
performance results from an inverse relationship between population
size and the mutation rate, i.e. high mutation rates were better for
small population sizes and low mutation rates were good for large
populations. Whilst recognizing that their results may not
generalize beyond the 10 functions in their test suite, they
recommend the following parameter values:
- Population size: 20 - 30
- Mutation rate: 0.005 - 0.1
- Cross-over rate: 0.75 - 0.95
De Jong's work was very important in that it provided practical
guidelines for subsequent applications of GA's. His recommendations
for the various parameters have been so widely adopted that they are
sometimes referred to as the "standard settings".
7.6.2 Theoretical Studies
Several researchers have theoretically analysed the dynamics of GAs.
In his survey, Lobo (2000: chapter 3) reports the most notable of
these as being the work on selection by Goldberg and Deb (1991); the
work on mutation by Mûhlenbein (1992) and Bäck (1993); the
research on population sizing by Goldberg, Deb and Clark (1992) and
Harik et al. (1997); and the work on control maps by Goldberg, Deb
and Thierens (1993) and Thierens and Goldberg (1993). The insights
and practical implications afforded by these studies are summarized
below.
- On Selection. In the absence of all other operators,
repeated use of the selection operator would eventually result in a
population comprised of the single chromosome with the highest
fitness. Goldberg and Deb define the takeover time as the
time it takes (as measured by the number of generations elapsed) for
this event to occur. They derived takeover time formulae for various
selection schemes and validated these using computer simulations.
For fitness proportionate selection schemes, the takeover time
depends on the fitness function distribution; for order-based
selection, the takeover time is independent of the fitness function
and is of the order O (log P), where P is the population size.
Obviously the takeover time is increases in the presence of
cross-over and mutation, but one must be careful not to exert too
much selection pressure to cancel the diversifying effects of these
operators.
- On Mutation. Independently of each other Mûhlenbein
(1992) and Bäck (1993) analyzed the effects of mutation on a
simple (1 + 1) evolutionary algorithm. They both concluded that for
a chromosome of length L, the optimal fixed mutation rate is
L−1. Intuitively, it is easy to see why there should be this
inverse relationship between chromosome length and mutation rate.
Besides exploring the search space, the mutation (and to so extent,
cross-over operation) can disrupt building blocks during the course
of a GA run. And obviously, this is more likely to occur for long
chromosomes than short ones since the operator is applied (with
probability) to each gene. So in order to minimize building block
disruption, one should decrease the mutation rate for relatively
longer chromosomes.
- On Population Size. Studies on population size attempt
to formulate schema growth equations similar to equation
5 that have population size as a variable. Unfortunately,
population sizing
equations are difficult to use in practice. Lobo notes:
"In order to apply [the equation] the user has to know or
estimate the selective advantage that a building block has over its
closest competitors; he has to estimate the building block's fitness
variance, he has to estimate the maximum level of deception in the
problem at hand; and of course he has to hope that the building
blocks are going to mix well, which may not occur in practice"
(paraphrased from p.34)
It is difficult to see how these population sizing studies can be
used to further inform the choice of parameter values suggested by
the empirical work of De Jong (1975) or Schaffer et al. (1989).
- On Selection. Increasing the population size resulted
in better long-term performance, but smaller population sizes
responded faster and therefore exhibited better initial performance.
7.6.3 Parameter Adaptation
We mention, in passing, parameter adaptation techniques. These
methods change GA parameters as the search progresses. There are
three main approaches: (1) centralized methods change parameter
values based on a central learning rule; (2) decentralized methods
encode the parameters values into the chromosomes themselves; (3)
meta-GA's attempt to optimize parameter values by evolving these
values for the actual GA that is run at a lower level using the
parameters identified by the meta-level GA. The main advantage of a
GA so designed is that the user is no longer required to specify
parameter values prior to executing the search.
7.7 Concluding Remarks
Genetic Algorithms are simple and yet powerful search and
optimization procedures that are widely applicable. Unfortunately,
our current knowledge is such that one cannot rigorously predict
whether a GA is going to efficient in any given instance due to the
difficult in choosing the parameters of the algorithm. Nevertheless,
the parameters recommended for GENO are efficient, at least on the
examples reported. These parameters were arrived at after extensive
"trial and error" experimentation guided by the empirical and
theoretical work outlined above: they are summarized in Section
3.1.
« Previous « Start
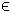 [x_L,
x_U]
[x_L,
x_U]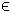 [x_L,
x_U]
[x_L,
x_U]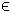 [x_L,x_U]
[x_L,x_U]